We don’t have a live CDR but I will see what we can do/if there’s an appatite for NWIS to jump on board. We have a means fo getting this pushed out quickly as a dedicated eform in the Welsh Clinical Portal so this could work.
J
Had a quick look. Was thinking about this over lunch with respect more to a patient self assessment tool in 1st instance but can see why you have steered away from that.
Have you had any chats with NDS about this? My thinking is that the blockers to any rollout and scale up would be staff authentication and patient ID at this stage for us, although of course Forth Valley have these integrations now so maybe an opportunity?
@Paulmiller - that was thinking exactly - great fit for FV, much harder for community or other settings.
Patient identity does not go away for us but by housing the app within WCP, we always maintain the “patient context” which comrises of NHS number and a varity of hospital numbers form the MPI. It might mean we inialise ehrId’s with hospital opposed to NHS numbers where these are not verified - not ideal but certinaly not a major problem either.
We could, if needed, just commit canoncial XML to our doc repo…  while we get a CDR up and running.
while we get a CDR up and running.
Have you spoken to them or anyone at NDS about this yet? Of course introducing a new form / tech / data access at a time of ‘emergency’ may just add to complexity of what they need to do so I guess it may not be a help! People will I expect want to work with the IT they currently have because, even if it is not very good, they know it works and how to use it.
Be worth highlighting though.
Sure - there may be some very practical barriers and pushback, esp as FV have only just gone live with ReSPECT but if nothing else we should have the discussion.
I guees one issue that might make the openEHR option more compelling is that within a hospital setting, this really is EHR data - it will need to be properly stored, versioned, audit logged because folks might be sued down the line. This is quite a different scenario to a citizen-facing tool where there is no datacapture.
The other thing is that we can change things very quickly and cheaply as/ when the epidemiology morphs.
Who would be best to chat to at NDS?
Ian
Alistair E / Johnny W probably. Use a Teams chat and cc me in.
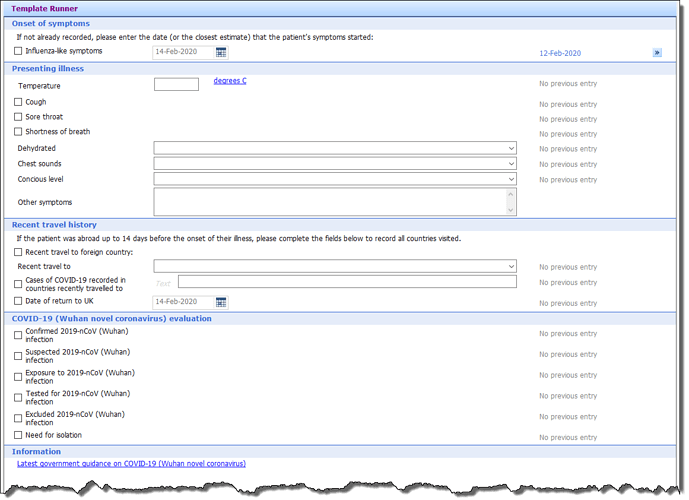
I try to write out some of the thinkings in the application we made. We named the product “On track of COVID-19” .
The document is work in progress and you find it here.
If you find it useful - use it. If you don’t - be happy. If you want to improve it - just edit.
Attached is some screenshots from the application we made. I am sorry the screenshots are in Norwegian. Given the “time to market” we didn’t focus on the multi-language support.
COVID-19-Tracker.pptx (2.0 MB)
NHS Digital will be releasing an emergency SNOMED Release via TRUD tomorrow, containing new codes relating to 2019 nCoV surveillance and management.
A release notification will be sent to the usual relevant subpack subscribers; the text of this notification will be as below:
Dear ,
An unscheduled emergency release of SNOMED CT UK Clinical Edition, RF2 for February 2020 is now available for download from the following links:
UK SNOMED CT Clinical Edition, RF2: Full, Snapshot & Delta download here
UK SNOMED CT Clinical Edition, RF2: Full, Snapshot & Delta download in “Beta” release file bundling here
UK SNOMED CT Clinical Edition, RF2: Delta download here
The SNOMED CT UK Drug extension is not released as part of this unscheduled SNOMED CT release. SNOMED CT UK Drug Extension data is published on a four-weekly cycle within the “UK Drug Extension, RF2” sub pack on TRUD. Please review the associated documentation for more details.
Release documentation is now only accessible on DELEN, at SNOMED CT UK Clinical Extension Release Documentation - Delen: Home - NHS Digital.
Information regarding this release:
This 28.1.1 release remains based on the July 2018 International Edition, whose release files are also included within the 28.1.1 release package.
This is a minor version release; content is identical to the existing 28.0.0 release of October 2019 except for the addition of the following codes, terms and synonyms in relation to the 2019 novel coronavirus:
Clinical finding
1240581000000104 2019-nCoV (novel coronavirus) detected
1240591000000102 2019-nCoV (novel coronavirus) not detected
1240631000000102 Did not attend 2019-nCoV (novel coronavirus) vaccination
1240751000000100 Disease caused by 2019-nCoV (novel coronavirus)
1240561000000108 Encephalopathy caused by 2019-nCoV (novel coronavirus)
1240571000000101 Gastroenteritis caused by 2019-nCoV (novel coronavirus)
1240601000000108 High priority for 2019-nCoV (novel coronavirus) vaccination
1240531000000103 Myocarditis caused by 2019-nCoV (novel coronavirus)
1240521000000100 Otitis media caused by 2019-nCoV (novel coronavirus)
1240551000000105 Pneumonia caused by 2019-nCoV (novel coronavirus)
1240541000000107 Upper respiratory tract infection caused by 2019-nCoV (novel coronavirus)
Event
1240431000000104 Exposure to 2019-nCoV (novel coronavirus) infection
Observable entity
1240741000000103 2019-nCoV (novel coronavirus) serology
Procedure
1240491000000103 2019-nCoV (novel coronavirus) vaccination
1240511000000106 Detection of 2019-nCoV (novel coronavirus) using polymerase chain reaction technique
1240461000000109 Measurement of 2019-nCoV (novel coronavirus) antibody
1240471000000102 Measurement of 2019-nCoV (novel coronavirus) antigen
1240451000000106 Telephone consultation for suspected 2019-nCoV (novel coronavirus)
Qualifier value
1240421000000101 Serotype 2019-nCoV (novel coronavirus)
Situation with explicit context
1240661000000107 2019-nCoV (novel coronavirus) vaccination contraindicated
1240651000000109 2019-nCoV (novel coronavirus) vaccination declined
1240781000000106 2019-nCoV (novel coronavirus) vaccination invitation short message service text message sent
1240681000000103 2019-nCoV (novel coronavirus) vaccination not done
1240671000000100 2019-nCoV (novel coronavirus) vaccination not indicated
1240701000000101 2019-nCoV (novel coronavirus) vaccine not available
1240731000000107 Advice given about 2019-nCoV (novel coronavirus) by telephone
1240721000000105 Advice given about 2019-nCoV (novel coronavirus) infection
1240711000000104 Educated about 2019-nCoV (novel coronavirus) infection
1240761000000102 Suspected disease caused by 2019-nCoV (novel coronavirus)
Substance
1240401000000105 Antibody to 2019-nCoV (novel coronavirus)
1240391000000107 Antigen of 2019-nCoV (novel coronavirus)
1240411000000107 Ribonucleic acid of 2019-nCoV (novel coronavirus)
The cross mapping component of this release is unchanged from the 28.0.0 data: it includes only the previously published maps to OPCS 4.8 and ICD10.
NB: Novel coronavirus codes in the Jan 2020 International Edition
A similar emergency update release of the January 2020 International Edition from SNOMED International occurred on January 31st, adding the following 2019 novel coronavirus codes:
840539006|Disease caused by 2019 novel coronavirus (disorder)|
840534001|2019 novel coronavirus vaccination (procedure)|
840544004|Suspected disease caused by 2019 novel coronavirus (situation)|
840536004|Antigen of 2019 novel coronavirus (substance)|
840535000|Antibody to 2019 novel coronavirus (substance)|
840533007|2019 novel coronavirus (organism)|
840546002|Exposure to 2019 novel coronavirus (event)|
However, this version of the International Edition can not currently be used within the UK because our current Clinical and Drug Extension releases are not compatible with it. The UK Edition as a whole (both Clinical, Drug and Pathology Extensions) is working to restore full alignment with prevailing current International Releases from October 2020. However, until that alignment and compatibility is restored, these seven International Edition codes should therefore not be used within the UK.
When alignment is achieved, and the new International coronavirus codes therefore become part of the UK Edition, the seven corresponding UK-only codes will be inactivated. Appropriate entries will be made in the der2_cRefset_Association tables, and so also in our own History Substitution and Query Table products derived from those, stating an equivalence between (for example) 840544004 and 1240761000000102.
Hi all,
I am adding 2 more files that could be useful for this topic
covid-19-patient-pathway-v2.3.pdf (758.4 KB)
WHO-2019-nCoV-SurveillanceGuidance-2020.4-eng.pdf (637.5 KB)
The latest WHO reporting spreadsheets are at
I have made a decent start analysing this and will upload the template ASAP
UK E-MIS GP system
These will almost all have associated SNOMED CT codes - I will try to find out which ones!!
Ian
openEHR-Suspected Covid-19 assessment.v0 (1).zip (279.7 KB)
openEHR-Suspected Covid-19 assessment.v0.oet (39.6 KB)
openEHR-Suspected Covid-19 assessment.v0.opt (707.3 KB)
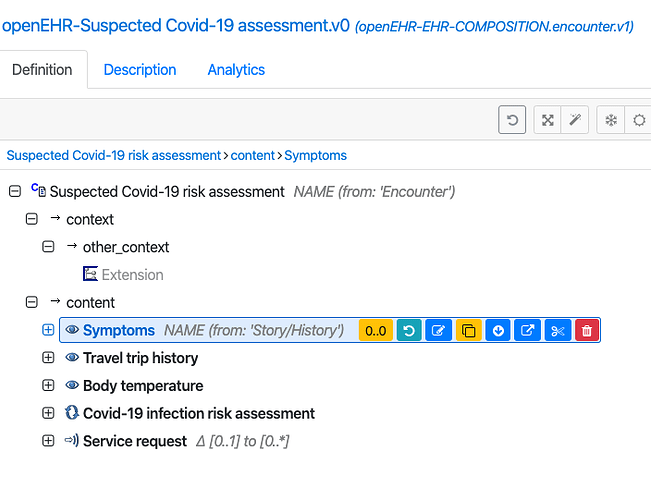
Considerably modified and tidied up Covid-19 screening template. I have tried to minimise any dependency on English - rermoved ad-hoc sections, added some specialisations, used symptom-sign specialisation, and removed the list of countries/regions so that this can be inserted or served in local languages.
I’ve had an issue uploading to CKM for some weird reason that I’m going to ask Sebastian Garde to look at ASAP.
Bedtime!!
@siljelb - there seeem to be a mistmatch on the Symptom arcehtype between Norwegian and openEHR CKM.
Norwegian: https://arketyper.no/ckm/archetypes/1078.36.358
ELEMENT[at0186] occurrences matches {0..1} matches { -- Første tilfelle?
value matches {
DV_BOOLEAN matches {
value matches {True}
}
}
}
openEHR CKM https://www.openehr.org/ckm/archetypes/1013.1.195
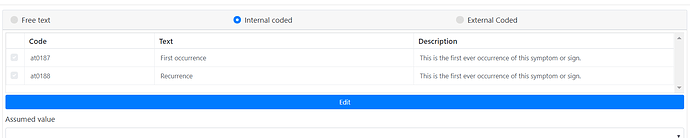
ELEMENT[at0186] occurrences matches {0..1} matches { -- Occurrence
value matches {
DV_CODED_TEXT matches {
defining_code matches {
[local::
at0187, -- First occurrence
at0188] -- Recurrence
}
}
}
}
Do you remember why?
@ian.mcnicoll - there is a minor bug in the symptom archetype on the occurences description.
It’s the same description for “First occurence” and “Recurrence”.
This is an as of yet unpublished (breaking) change in the international archetype that hasn’t been propagated to the Norwegian CKM yet.
Thanks.
Use of the breaking change at own risk. That’s fine for an application related to “risk management” - infection risk